Introduction to Bayesian Optimization (BO)¶
Bayesian optimization is a model-based, black-box optimization algorithm that is tailored for very expensive objective functions (a.k.a. cost functions) [1][3]. As a black-box optimization algorithm, Bayesian optimization searches for the maximum of an unknown objective function from which samples can be obtained (e.g., by measuring the performance of a robot). Like all model-based optimization algorithms (e.g. surrogate-based algorithms, kriging, or DACE), Bayesian optimization creates a model of the objective function with a regression method, uses this model to select the next point to acquire, then updates the model, etc. It is called Bayesian because, in its general formulation [3], this algorithm chooses the next point by computing a posterior distribution of the objective function using the likelihood of the data already acquired and a prior on the type of function.
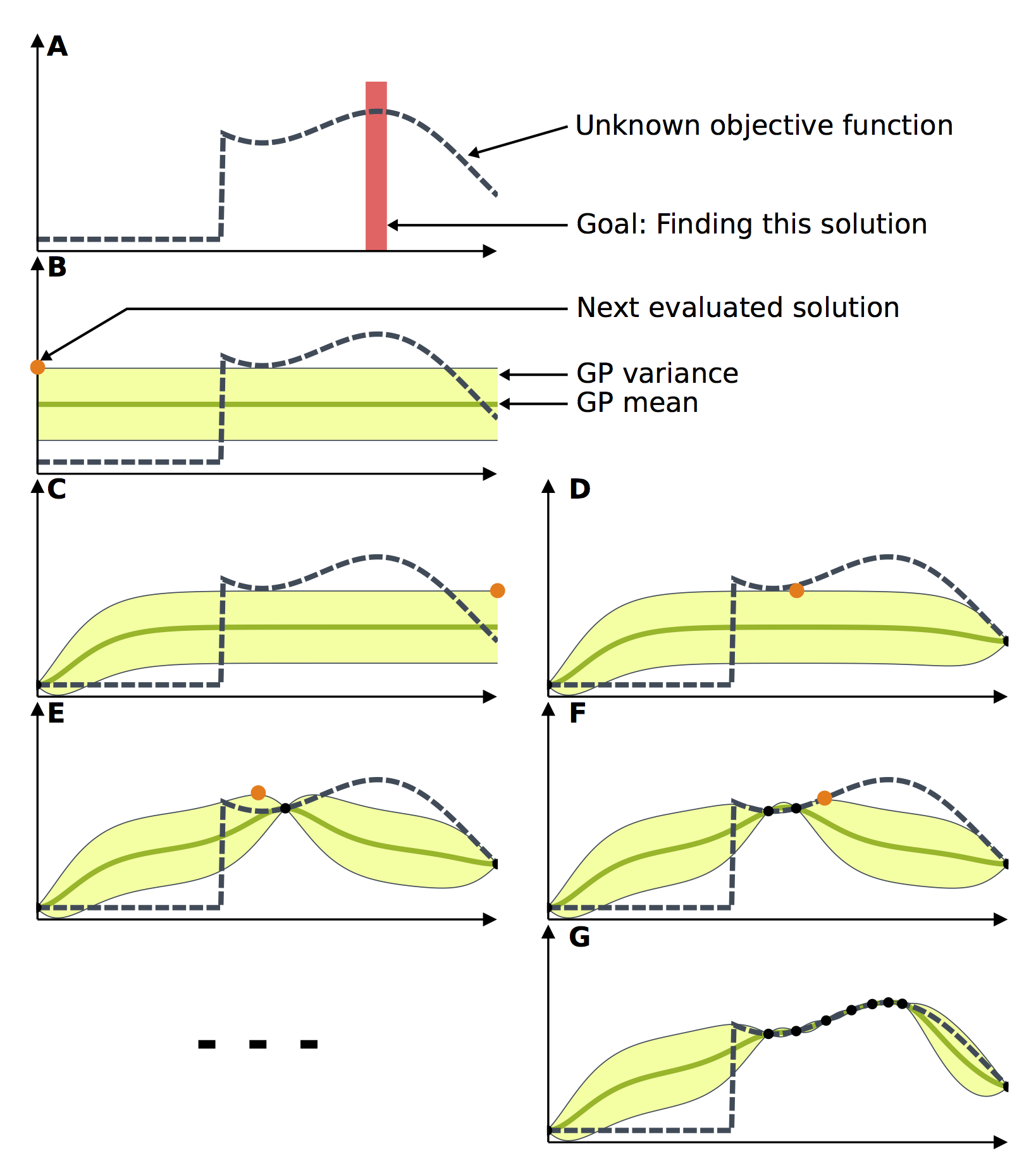
Bayesian Optimization of a toy problem. (A) The goal of this toy problem is to find the maximum of the unknown objective function. (B) The Gaussian process is initialized, as it is customary, with a constant mean and a constant variance. (C) The next potential solution is selected and evaluated. The model is then updated according to the acquired data. (D) Based on the new model, another potential solution is selected and evaluated. (E-G) This process repeats until the maximum is reached.
Gaussian Process¶
Limbo uses Gaussian process regression to find a model [4], which is a common choice for Bayesian optimization [1]. Gaussian processes are particularly interesting for regression because they not only model the cost function, but also the uncertainty associated with each prediction. For a cost function \(f\), usually unknown, a Gaussian process defines the probability distribution of the possible values \(f(\mathbf{x})\) for each point \(\mathbf{x}\). These probability distributions are Gaussian, and are therefore defined by a mean (\(\mu\)) and a standard deviation (\(\sigma\)). However, \(\mu\) and \(\sigma\) can be different for each \(\mathbf{x}\); we therefore define a probability distribution over functions:
where \(\mathcal{N}\) denotes the standard normal distribution.
To estimate \(\mu(\mathbf{x})\) and \(\sigma(\mathbf{x})\), we need to fit the Gaussian process to the data. To do so, we assume that each observation \(f(\mathbf{\chi})\) is a sample from a normal distribution. If we have a data set made of several observations, that is, \(f(\mathbf{\chi}_1), f(\mathbf{\chi}_2), ..., f(\mathbf{\chi}_t)\), then the vector \(\left[f(\mathbf{\chi}_1), f(\mathbf{\chi}_2), ..., f(\mathbf{\chi}_t)\right]\) is a sample from a multivariate normal distribution, which is defined by a mean vector and a covariance matrix. A Gaussian process is therefore a generalization of a \(n\)-variate normal distribution, where \(n\) is the number of observations. The covariance matrix is what relates one observation to another: two observations that correspond to nearby values of \(\chi_1\) and \(\chi_2\) are likely to be correlated (this is a prior assumption based on the fact that functions tend to be smooth, and is injected into the algorithm via a prior on the likelihood of functions), two observations that correspond to distant values of \(\chi_1\) and \(\chi_2\) should not influence each other (i.e. their distributions are not correlated). Put differently, the covariance matrix represents that distant samples are almost uncorrelated and nearby samples are strongly correlated. This covariance matrix is defined via a kernel function, called \(k(\chi_1, \chi_2)\), which is usually based on the Euclidean distance between \(\chi_1\) and \(\chi_2\) (see the “kernel function” sub-section below).
Given a set of observations \(\mathbf{P}_{1:t}=f(\mathbf{\chi}_{1:t})\) and a sampling noise \(\sigma^2_{noise}\) (which is a user-specified parameter), the Gaussian process is computed as follows [1][4]:
Our implementation of Bayesian optimization uses this Gaussian process model to search for the maximum of the unknown objective function \(f(\mathbf{x})\). It selects the next \(\chi\) to test by selecting the maximum of the acquisition function, which balances exploration – improving the model in the less explored parts of the search space – and exploitation – favoring parts that the models predicts as promising. Once an observation is made, the algorithm updates the Gaussian process to take the new data into account. In classic Bayesian optimization, the Gaussian process is initialized with a constant mean because it is assumed that all the points of the search space are equally likely to be good. The model is progressively refined after each observation.
Optimizing the hyper-parameters of a Gaussian process¶
A GP is fully specified by its mean function \(\mu(\mathbf{x})\) and covariance function \(k(\chi_1, \chi_2)\) (a.k.a. kernell function). Nevertheless, the kernel function often includes some parameters, called hyperparameters, that need to be tuned. For instance, one of the most common kernel is the Squared Exponential covariance function:
For some datasets, it makes sense to hand-tune these parameters (e.g., when there are very few samples). Ideally, our objective should be to learn \(l^2\) (characteristic length scale) and \(\sigma_f^2\) (overall variance).
A classic way to do so is to maximize the probability of the data given the hyper-parameters \(\mathbf{\vartheta}\) (there are other ways, e.g. cross-validation). We use a log because it makes the optimization simpler and does not change the result.
The marginal likelihood can be computed as follows:
where \(\mu_0\) is the mean function (prior).
Limbo provides many algorithms to optimize the likelihood. Some algorithms are gradient-free (e.g. CMA-ES), some others use the gradient of the log-likelihood (e.g. rprop), see Optimization Sub-API and the the Limbo implementation guide.
For more details, see [4] (chapter 5).
Kernel function¶
The kernel function is the covariance function of the Gaussian process. It defines the influence of a solution’s performance on the performance and confidence estimations of not-yet-tested solutions that are nearby.
The Squared Exponential covariance function and the Matern kernel are the most common kernels for Gaussian processes [1][4]. Both kernels are variants of the “bell curve”. The Matern kernel is more general (it includes the Squared Exponential function as a special case) and allows us to control not only the distance at which effects become nearly zero (as a function of parameter \(\rho\)), but also the rate at which distance effects decrease (as a function of parameter \(\nu\)).
The Matern kernel function is computed as follows [2][5] (with \(\nu=5/2\)):
There are other kernel functions in Limbo, and it is easy to define more. See the Limbo implementation guide for the available kernel functions.
Acquisition function¶
In order to find the next point to evaluate, we optimize the acquisition function over the model. This step is another optimization problem, but does not require evaluating the objective function. In general, for this optimization problem we can derive the exact equation and find a solution with gradient-based optimization, or use any other optimizer (e.g. CMA-ES).
Several different acquisition functions exist, such as the probability of improvement, the expected improvement, or the Upper Confidence Bound (UCB) [1]. For instance, the equation for the UCB is:
where \(\kappa\) is a user-defined parameter that tunes the tradeoff between exploration and exploitation.
Here, the emphasis on exploitation vs. exploration is explicit and easy to adjust. The UCB function can be seen as the maximum value (argmax) across all solutions of the weighted sum of the expected performance (mean of the Gaussian, \(\mu_{t}(\mathbf{x})\)) and of the uncertainty (standard deviation of the Gaussian, \(\sigma_t(\mathbf{x})\)) of each solution. This sum is weighted by the \(\kappa\) factor. With a low \(\kappa\), the algorithm will choose solutions that are expected to be high-performing. Conversely, with a high \(\kappa\), the algorithm will focus its search on unexplored areas of the search space that may have high-performing solutions. The \(\kappa\) factor enables fine adjustments to the exploitation/exploration trade-off of the algorithm.
There are other acquisition functions in Limbo, and it is easy to define more. See the Limbo implementation guide for the available acquisition functions.
| [1] | (1, 2, 3, 4, 5) Eric Brochu, Vlad M Cora, and Nando De Freitas. A tutorial on bayesian optimization of expensive cost functions, with application to active user modeling and hierarchical reinforcement learning. arXiv preprint arXiv:1012.2599, 2010. |
| [2] | Bertil Matérn and others. Spatial variation. stochastic models and their application to some problems in forest surveys and other sampling investigations. Meddelanden fran statens Skogsforskningsinstitut, 1960. |
| [3] | (1, 2) J. Mockus. Bayesian approach to global optimization: theory and applications. Kluwer Academic, 2013. |
| [4] | (1, 2, 3, 4) C. E. Rasmussen and C. K. I. Williams. Gaussian processes for machine learning. MIT Press, 2006. ISBN 0-262-18253-X. |
| [5] | Michael L Stein. Interpolation of spatial data: some theory for kriging. Springer, 1999. |